
the hyperplane parameter values to those found during the fitting process. support_vectors_ yy_up = a * xx + ( b - a * b ) # plot the line, the points, and the nearest vectors to the plane pl. These functions produce helpful 2d and 3d diagnostic plots for post hyper.fit. support_vectors_ yy_down = a * xx + ( b - a * b ) b = clf. intercept_ ) / w # plot the parallels to the separating hyperplane that pass through the # support vectors b = clf. linspace ( - 5, 5 ) yy = a * xx - ( clf.

You should not call this function directly, only through the method. Note This function is available as the plot () method of hyperplane arrangements. If the arrangement is in 4 dimensions but inessential, a plot of the essentialization is returned. fit ( X, Y ) # get the separating hyperplane w = clf. Return a plot of the hyperplane arrangement. randn ( 20, 2 ) + ] Y = * 20 + * 20 # fit the model clf = svm.
PLOT HYPERPLAN HOW TO
I'm not sure how to plot a number line, but you can always resort to a scatter plot with all y coordinates set to 0.Print _doc_ import numpy as np import pylab as pl from sklearn import svm # we create 40 separable points np. So to visualize, you merely need to plot your data on a number line (use different colours for the classes), and then plot the boundary obtained by dividing the negative intercept by the slope. equate the last entry in w as the hyperplane offset b and write the separating hyperplane equation y wx+b with w ( 1. The surface is made opaque by using antialiasedFalse.
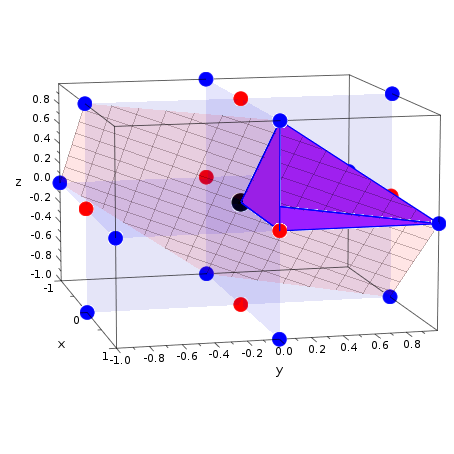
That's your 0-d hyperplane (point) for the classifier. Demonstrates plotting a 3D surface colored with the coolwarm colormap. A simulated SAR plot for the central cross-section is then displayed on the. Here are plots of 1st versus 2nd, 3rd versus 4th, and 5th versus 6th. Now try changing the means of the distributions that make up X, and you will find that the answer is always -intercept/slope. Since Sigma-HyperPlan does not account for the convective nature of heat. Principal Component Analysis (PCA) is one way of finding a hyperplane that is.
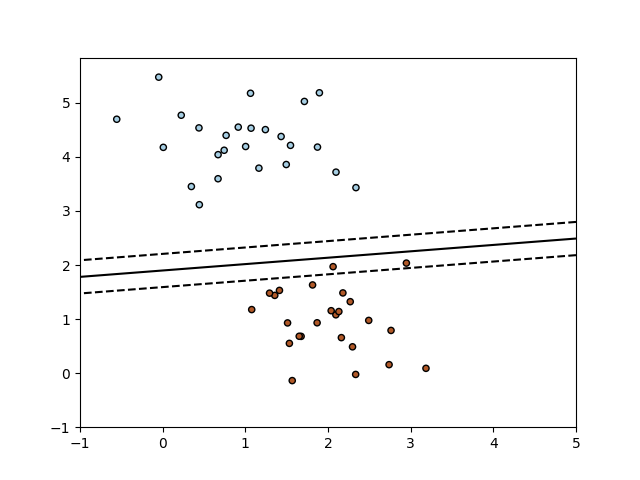
However, if you take y=0 and back-calculate x, it will be pretty close to 5. In geometry, a hyperplane of an n-dimensional space V is a subspace of dimension n 1, or equivalently, of codimension 1 in V.The space V may be a Euclidean space or more generally an affine space, or a vector space or a projective space, and the notion of hyperplane varies correspondingly since the definition of subspace differs in these settings in all cases. Once you fit the classifier, you find out the intercept is about -0.96 which is nowhere near where the 0-d hyperplane (i.e. Intuitively it's clear the hyperplane should be halfway between 0 and 10. y is the array of labels (100 of class '0' and 100 of class '1'). Two-dimensional partial dependence plot showing the.
PLOT HYPERPLAN DOWNLOAD
X is the array of samples such that the first 100 points are sampled from N(0,0.1), and the next 100 points are sampled from N(10,0.1). Download scientific diagram 2: A good separating hyperplane is an hyperplane that maximizes the.
PLOT HYPERPLAN CODE
So I decided to play with the sample code below to see if I can figure out the answer: from sklearn import svm I've spent about an hour looking for answers in the documentation for scikit-learn, but there is simply nothing on 1-d SVM classifiers (probably because they are not practical). So the question is really how to turn this line into a point. Yet what scikit-learn gives you is a line. On the surface it's very simple - one feature means one dimension, hence the hyperplane has to be 0-dimensional, i.e. You can find the coefficients (and ) using the two equations below.Quoting from 'Support-Vector Networks' by Cortes and Vapnik, 1995, '.
